|
5/22/2023 0 Comments Medinria update 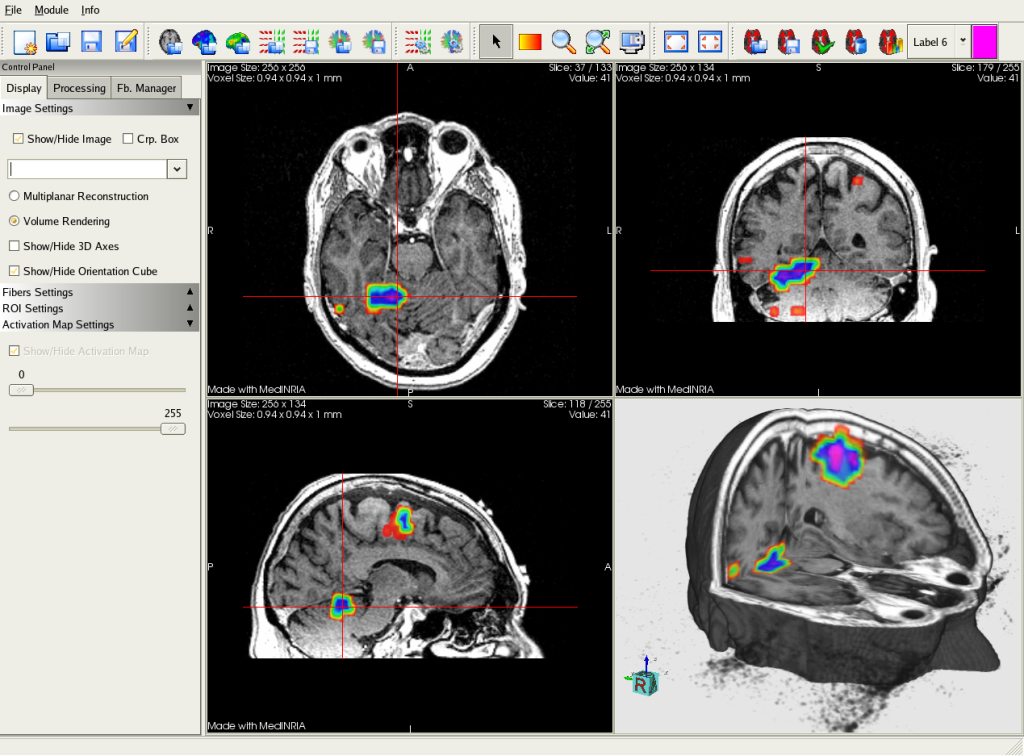
All VTK meshes and line sets are also supported and may be viewed on top of other images, providing a comprehensive way e.g. We also provide the usual slice by slice navigation as well as 3D/4D volume rendering of images. You may view typical 2D/3D images, side by side or superimposed on separate layers. MedInria provides a powerful environment to visualize many kinds of medical data. Visualization: 2D to 4D image visualization, tensor, surfaces visualization in a drag-drop and tabs interface.Finally, it also allows for a quick preview of data in your database through thumbnails. It is able to load any format that is supported by ITK and GIS images. It provides the usual features related to data management: import or index data from various data sources (file system, Dicom, external databases - Shanoir, PACS server), export data to disk. MedInria comes with a patient centric database, accessible in the Browser. Database management and file import: DICOM read, PACS interrogation, various file formats handling, etc.It provides in its current release 6 main feature categories including: MedInria can be used for a wide variety of tasks. Through an intuitive user interface, medInria offers from standard to cutting-edge processing functionalities for your medical images such as 2D/3D/4D image visualization, image registration, diffusion MR processing and tractography. I think there is still an argument that Napari could provide similar functionality, but in a less domain-specific manner and with the flexibility to combine interactively with other Python image processing tools.MedInria is a multi-platform medical image processing and visualization software. I think the main argument against developing this is that there are already good non-Python tools that do this. Often there is an (optional) crosshair overlaid on each view to show the current location across views. PitchĪ key feature is that clicking on a specific voxel in one view, updates the other two orthogonal views to show the slice at the selected location. Nibabel provides a basic 3D orthogonal viewer built using Matplotlib, but Napari could potentially produce something much nicer. I saw that you can already switch between the orthogonal views in the main window, but it would be nice to be able to see them side by side and have the spatial positions linked across the views. Perhaps this is already possible today? I have only just started looking at napari. 
similar to this screenshot from 3D slicer, although as a first step it could exclude the 4th panel with the 3D rendering)? Another option sometimes used for a 4th panel is to show the time-course of the selected 3D spatial position when the data is 3D + time. I found the following duplicate issue in an archived repository: napari/napari-core#4,ĭo you know if there is anyone working on a plugin that would show simultaneous orthogonal views of 3D data (e.g. 
It is also great that this viewer can be called directly on general NumPy and Dask arrays as opposed to requiring specific medical imaging file formats. freeview, FSLeyes, 3D Slicer, Medinria, etc.), but I am excited to see something in the Python space. I think the only mouse-control I am used to having in other applications that I didn't see was the ability to adjust contrast and window level by dragging the mouse in combination with one of the mouse buttons (I did see there is a slider for it in the left panel).Īs you may be aware, there are quite a few good standalone viewers/tools developed in C++ for medical imaging applications (e.g. I tried it out yesterday on some large (~16GB) 3D data and browsing through slices was very smooth! Sliding and zooming the volume worked well. I work in medical imaging and often collaborate with on scikit-image, but for some reason had not gotten around to trying napari. I first want to congratulate the napari devs for their great work on this package. 
0 Comments
Leave a Reply. |

 RSS Feed
RSS Feed
